The strange thing is that this cool discovery went nowhere even though it would have some really interesting applications. In fact, this paper was followed by two others, then a hiatus and then two papers in the 90s about melting temperature. No cloning, no nothing.
The phage was lost as there is a patent that gives its sequence. The sequence is unhelpfully unannounced, but the patent among the really boring bits has claim 270, which says:
On the other hand, it seems very likely that the D-base is formed by semi-replicative modification. Between the two biosynthesis routes of dDTP formation described above, the identification of a succinyladenylate synthetase gene homologue called ddbA (deoxyribodiaminopurine biosynthetic gene A) leads to the conclusion that it is the second route which is probably taken during phage infection (FIG. 2).
Several tests have been carried out in order to determine the activity of the corresponding protein. The results suggest that the expression of ddbA allows restoration of the growth of a strain of E. coli expressing the yaaG gene of Bacillus subtilis [yaaG (now dgk) encodes deoxyguanosine kinase] in the presence of a high concentration of dG (10 mM). On the other hand, 2,6-diaminopurine becomes toxic (10 mM) to E. coli when it is in phosphorylated form (which has been tested in the same strain of E. coli expressing the yaaG gene of Bacillus subtilis i.e. MG1655 pSU yaaG) which makes it possible to have a screen in order to identify in vivo the complete biosynthesis route of the D-base.
Several tests have been carried out in order to determine the activity of the corresponding protein. The results suggest that the expression of ddbA allows restoration of the growth of a strain of E. coli expressing the yaaG gene of Bacillus subtilis [yaaG (now dgk) encodes deoxyguanosine kinase] in the presence of a high concentration of dG (10 mM). On the other hand, 2,6-diaminopurine becomes toxic (10 mM) to E. coli when it is in phosphorylated form (which has been tested in the same strain of E. coli expressing the yaaG gene of Bacillus subtilis i.e. MG1655 pSU yaaG) which makes it possible to have a screen in order to identify in vivo the complete biosynthesis route of the D-base.
- either aminated to adenine in two steps via a N-succinyl intermediate by the enzymes encoded by purA and purB
- or oxidised at position 2 (resulting in a keto substituted compound, xanthosine monophosphate) and transaminated to guanine.
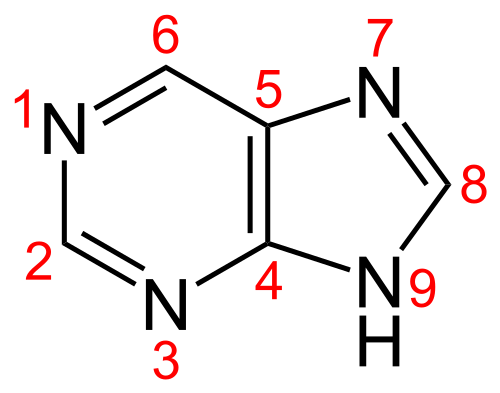
Consequently, to make diaminopurine monophosphate the cyanophage would need to aminate position 6 of guanosine monophosphate (guanine has a ketone at 6 and an amine at 2).
The authors claim that there is a homologue of the purA, called ddbA, whose enzyme can act on GMP, unlike the PurA enzyme.
However, they do not report a purB homologue or a adk homologue (encoding a kinase to make the triphosphate). This means that:
- either Synechococcus (the host) purB and adk promiscously do the reactions, which seems odd given that the phage would give them a negative selection,
- or that an analogue eluded them manually annotating the genome 1970s style.
Parenthetically, the people that filed the patent were not the people who found the virus and no paper was published about it even though the first author of the patent studies odd nucleobases.
One interesting thing from the genome is the presence of many polymerases and of DNA gyrase, which is due to the odd DNA.
So why did nothing come out of it? Is it real?
There are four facts worth noting:
- Recently a paper came out in PNAS where they describe modified uridines in some phages. The concentration of modified uridine is a small percentage —which may likely be the case with this cyanophage too.
- Sanger sequencing or Illumina of a virus with diaminopurine will work just fine, except with Nanopore sequences, where the more charged base will have a shorter dwell time and result in a lot of failed reads.
- There were many papers about oxygen-insensitive nitrogenases in the 90s and 00s from a single team. It took a consortium of authors to disprove it thoroughly a decade later. In that case, there was a lot of secrecy about the strains and the genome of the organism. Whereas with cyanophage S-2L the sequence is on NCBI.
- That the S-2L strain is lost is suspicious, but not implausible especially how hard it is to grow cyanobacteria.
In conclusion, it is a real shame research into this curious pathway was minimal and has not progressed as it would be a perfect toolkit for synthetic biology.


No comments:
Post a Comment