Crosslinked residues are common, but for sure make up for it by being simultaneously highly intriguing and highly
technically problematic. Oddly, I seem to keep bumping into them. During my PhD a decade ago I saw a talk by the father
of Kiwi structural biochemistry, Ted Baker, about a curious case where they found an isopeptide bonds hidden in their
crystal density. In a postdoc I worked with isopeptide bonds
—I
blogged about isopeptide bonds in Rosetta
four years ago. During the start of the pandemic I dis some covalent-docking of compounds with PyRosetta for
the
Covid Moonshoot project, which evolved
into
Fragmenstein. Most tools have a hard time with crosslinks. And last
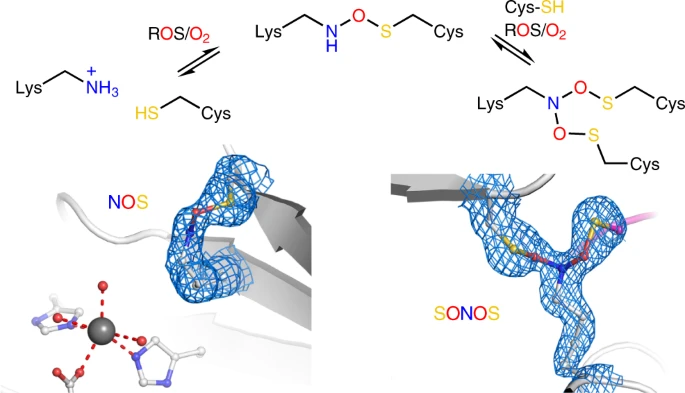
month the Twittersphere was abuzz with the news of lysine-hydroxylcysteine (N-O-S) bridges in protein.
PyMOL will strip LINK entries from PDBs on saving while NGL obeys only CONECT entries in PBDs. An exception is
PyRosetta: it behaves very nicely with disulfides, isopeptide bonds (
cf. repo of PyRosetta code
from Keeble et al.) and other crosslinks —mostly. As a
result I thought I'd add a note on how to add them in PyRosetta.